A Billion-Year Evolutionary Tale – Biologists Trace Cell Division Back to Its Roots
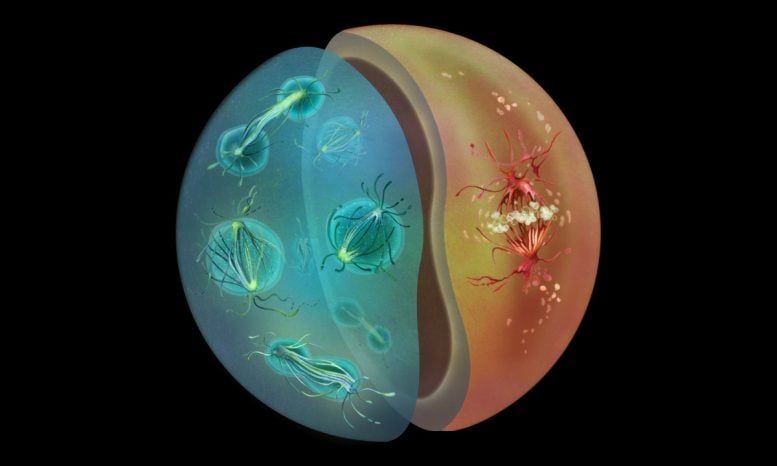
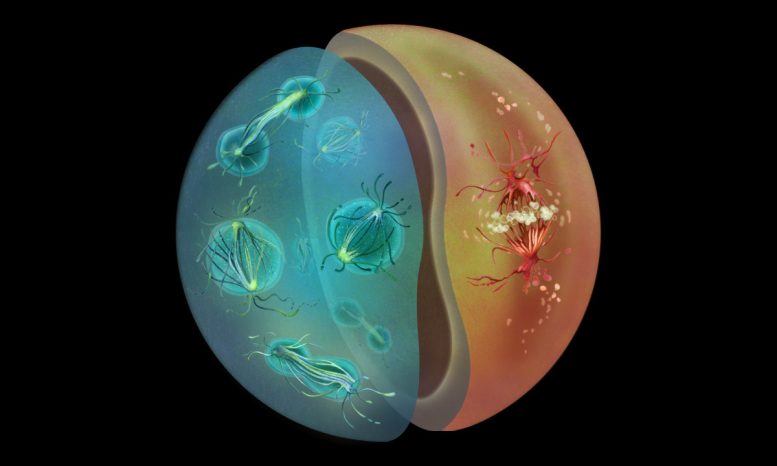
Ichthyosporeans Sphaeroforma arctica and Chromosphaera perkinsii undergoing mitosis, depicted as two halves of a cell, rendered in Haeckel-inspired tones and a naturalist style. Credit: Nirupa Rao
New findings by EMBL researchers reveal how animals and fungi have developed distinct cell division processes to accommodate their varied life cycles.
Cell division is a crucial process for all life forms, from bacteria to blue whales, enabling growth, reproduction, and the continuation of species. Despite its universal nature, the methods of cell division vary significantly across organisms. A recent study by EMBL Heidelberg’s Dey group, along with their collaborators and published in Nature, investigates the evolution of cell division methods in organisms closely related to fungi and animals. For the first time, this research demonstrates the connection between an organism’s life cycle and its cell division techniques.
Despite last sharing a common ancestor over a billion years ago, animals and fungi are similar in many ways. Both belong to a broader group called ‘eukaryotes’ – organisms whose cells store their genetic material inside a closed compartment called the ‘nucleus’. The two differ, however, in how they carry out many physiological processes, including the most common type of cell division – mitosis.
Most animal cells undergo ‘open’ mitosis, in which the nuclear envelope – the two-layered membrane separating the nucleus from the rest of the cell – breaks down when cell division begins. However, most fungi use a different form of cell division – called ‘closed’ mitosis – in which the nuclear envelope remains intact throughout the division process. However, very little is known about why or how these two distinct modes of cell division evolved and what factors determine which mode would be predominantly followed by a particular species.
Research Collaboration and Methodology
This question captured the attention of scientists in the Dey Group at EMBL Heidelberg, who investigated the evolutionary origins of the nucleus and cell division. “By studying diversity across organisms and reconstructing how things evolved, we can begin to ask if there are universal rules that underlie how such fundamental biological processes work,” said Gautam Dey, Group Leader at EMBL Heidelberg.
In 2020, during the COVID-19 lockdown, an unexpected path to answering this question grew out of discussions between Dey’s group and Omaya Dudin’s team at the Swiss Federal Institute of Technology (EPFL), Lausanne. Dudin is an expert in an unusual group of marine protists – Ichthyosporea. Ichthyosporea are closely related to both fungi and animals, with different species lying closer to one or the other group on the evolutionary family tree.
The Dey and Dudin groups, in collaboration with Yannick Schwab’s group at EMBL Heidelberg, decided to probe the origins of open and closed mitosis using Ichthyosporea as a model. Interestingly, the researchers found that certain species of Ichthyosporea undergo closed mitosis while others undergo open mitosis. Therefore, by comparing and contrasting their biology, they could obtain insights into how organisms adapt to and use these two cell division modes.
Hiral Shah, an EIPOD fellow working across the three groups, led the study. “Having recognized very early that Ichthyosporea, with their many nuclei and key evolutionary position between animal and fungi, were well-suited for addressing this question, it was clear that this would require bringing together the cell biological and technical expertise of the Dey, Dudin, and Schwab groups, and this is exactly what the EIPOD fellowship allowed me to do,” said Shah.
Upon closely probing the mechanisms of cell division in two species of Ichthyosporeans, the researchers found that one species, S. arctica, favors closed mitosis, similar to fungi. S. arctica also has a life cycle with a multinucleate stage, where many nuclei exist within the same cell – another feature shared with many fungal species as well as the embryonic stages of certain animals, such as fruit flies. Another species, C. perkinsii, turned out to be much more animal-like, relying on open mitosis. Its life cycle involves primarily mononucleate stages, where each cell has a single nucleus.
Implications for Understanding Eukaryotic Cell Division
“Our findings led to the key inference that the way animal cells do mitosis evolved hundreds of millions of years before animals did. The work therefore has direct implications for our general understanding of how eukaryotic cell division mechanisms evolve and diversify in the context of diverse life cycles, and provides a key piece of the animal origins puzzle,” said Dey.
The study combined expertise in comparative phylogenetics, electron microscopy (from the Schwab Group and the electron microscopy core facility (EMCF) at EMBL Heidelberg), and ultrastructure expansion microscopy, a technique that involves embedding biological samples in a transparent gel and physically expanding it. Additionally, Eelco Tromer, from the University of Groningen in the Netherlands, and Iva Tolic, from the Ruđer Bošković Institute in Zagreb, Croatia, provided expertise in comparative genomics and mitotic spindle geometry and biophysics, respectively.
“The first time we saw an expanded S. arctica nucleus, we knew this technique would change the way we study the cell biology of non-model organisms,” said Shah, who brought back the expansion microscopy technique to EMBL Heidelberg after a stint at the Dudin lab. Dey agrees: “A key breakthrough in this study came with our application of ultrastructure expansion microscopy (U-ExM) to the analysis of the ichthyosporean cytoskeleton. Without U-ExM, immunofluorescence and most dye labeling protocols do not work in this understudied group of marine holozoans.”
This study also demonstrates the importance of going beyond traditional model organism research when trying to answer broad biological questions, and the potential insights further research on Ichthyosporean systems might reveal. “Ichthyosporean development displays remarkable diversity,” said Dudin. “On one hand, several species exhibit developmental patterns similar to those of early insect embryos, featuring multinucleated stages and synchronized cellularisation. On the other hand, C. perkinsii undergoes cleavage division, symmetry breaking, and forms multicellular colonies with distinct cell types, similar to the ‘canonical view’ of early animal embryos. This diversity not only helps in understanding the path to animals but also offers a fascinating opportunity for comparative embryology outside of animals, which is, in itself, very exciting.”
The project’s inherent interdisciplinarity served not only as a good testbed for this type of collaborative research but also for the unique postdoctoral training afforded at EMBL. “Hiral’s project nicely illustrates the virtue of the EIPOD program: a truly interdisciplinary project, bundling innovative biology with advanced methods, all contributing to a truly spectacular personal development,” said Schwab. “We (as mentors) witnessed the birth of a strong scientist, and this is really rewarding!”
The Dey, Dudin, and Schwab groups are currently also collaborating on the PlanExM project, part of the TREC expedition – an EMBL-led initiative to explore and sample the biodiversity along European coasts. PlanExM aims to apply expansion microscopy to study the ultrastructural diversity of marine protists directly in environmental samples. “The project grew out of the realization that U-ExM is going to be a game-changer for protistology and marine microbiology,” said Dey. With this project, as well as others currently underway, the research team hopes to shed further light on the diversity of life on Earth and the evolution of the fundamental biological processes.
Reference: “Life-cycle-coupled evolution of mitosis in close relatives of animals” by Hiral Shah, Marine Olivetta, Chandni Bhickta, Paolo Ronchi, Monika Trupinić, Eelco C. Tromer, Iva M. Tolić, Yannick Schwab, Omaya Dudin and Gautam Dey, 22 May 2024, Nature.
DOI: 10.1038/s41586-024-07430-z


